Articles
- Page Path
- HOME > Endocrinol Metab > Volume 34(4); 2019 > Article
-
Review ArticleMedical Big Data Is Not Yet Available: Why We Need Realism Rather than Exaggeration
-
Hun-Sung Kim1,2
 , Dai-Jin Kim1,3, Kun-Ho Yoon1,2
, Dai-Jin Kim1,3, Kun-Ho Yoon1,2 -
Endocrinology and Metabolism 2019;34(4):349-354.
DOI: https://doi.org/10.3803/EnM.2019.34.4.349
Published online: December 23, 2019
1Department of Medical Informatics, College of Medicine, The Catholic University of Korea, Seoul, Korea.
2Department of Endocrinology and Metabolism, College of Medicine, The Catholic University of Korea, Seoul, Korea.
3Department of Psychiatry, College of Medicine, The Catholic University of Korea, Seoul, Korea.
- Corresponding author: Hun-Sung Kim. Department of Medical Informatics, College of Medicine, The Catholic University of Korea, 222 Banpo-daero, Seocho-gu, Seoul 06591, Korea. Tel: +82-2-2258-8262, Fax: +82-2-2258-8297, 01cadiz@hanmail.net
Copyright © 2019 Korean Endocrine Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
ABSTRACT
- Most people are now familiar with the concepts of big data, deep learning, machine learning, and artificial intelligence (AI) and have a vague expectation that AI using medical big data can be used to improve the quality of medical care. However, the expectation that big data could change the field of medicine is inconsistent with the current reality. The clinical meaningfulness of the results of research using medical big data needs to be examined. Medical staff needs to be clear about the purpose of AI that utilizes medical big data and to focus on the quality of this data, rather than the quantity. Further, medical professionals should understand the necessary precautions for using medical big data, as well as its advantages. No doubt that someday, medical big data will play an essential role in healthcare; however, at present, it seems too early to actively use it in clinical practice. The field continues to work toward developing medical big data and making it appropriate for healthcare. Researchers should continue to engage in empirical research to ensure that appropriate processes are in place to empirically evaluate the results of its use in healthcare.
- Artificial intelligence (AI) is one of the biggest talking points in the medical field today [123]. Big data, machine learning, deep learning, and the common data model (CDM), among others, are also commonly mentioned in the medical field [45678]. In fact, most medical staff wrongfully assume that medical big data can be easily extracted from electronic medical records (EMR), and analyzed using advanced statistics—a naïve assumption driven by the proliferation of medical data. In reality, extracting good data to use for research is not an easy task [45]. This is due to the inherent limitations of big data, specifically claim data, that had not been collected for research purposes. Ultimately, big data is a primitive form of data that needs to be manually reviewed for the extraction of useful and meaningful knowledge. Medical staff should be able to extract correct medical information in an environment overflowing with uncertain information.
- Limitations of machine learning
- Machine learning (or deep learning) is used to develop AI algorithms, and requires a large amount of data for full functionality. According to some scholars [910], machine learning performance increases along with the amount of data; however, the sophistication of analytic models does not improve performance significantly. In one study, an outstanding prediction model was developed using simple logistic regression [11]. Here, we are faced with a very basic question: why is deep learning necessary? In some cases, if the data is well refined, simple statistical procedures, like simple logistic regression, provide excellent results or prediction models, even without the application of complicated deep learning techniques that takes time to acquire, and requires high-end equipment. Due to the increased attention on machine learning [1213], methodological know-how is often perceived as the key for unlocking knowledge; however, the most important consideration in research is not the methodology, but rather how well the data extracted from the sample reflects the realities of the whole population. The use of complicated procedures and the development of statistical hypotheses reflect the fact that data quality cannot be guaranteed; when data is collected in a manner that reflects the unique qualities of the sample population, even simple statistics can produce sufficiently good results in some cases. This is why we continually emphasize the importance of data quality, rather than quantity. Of course, regardless of data quality, effective analysis may not be possible using traditional statistical methods, depending on the type of data. When digital data are standardized to a certain level and are well-managed in terms of quality (image or bio-signal data), existing traditional analytical methods cannot be used to elicit new interpretations. In such cases machine learning can be useful, despite limitations derived from inaccurate data or an absence of data quality management (DQM). Of great importance, however, is that medical staff are trained to look at data itself first to determine whether machine learning or traditional statistical methods would better serve the analytic purpose.
- Most clinical data contain noise and missing values, necessitating repetitive and labor-intensive tasks that involve time and effort, including data handling, DQM, and data cleansing [4]. Machine learning relies on data, and the deep learning itself is less important than the quality of data. If the data is unrefined, AI will not produce any good results, as exemplified by Microsoft's Tay Chatbot [14], which learned nonsense when it was taught nonsense. Various retrospective studies further demonstrated how undesirable phenomena produced different results, depending on the type of data and the research method [1516]. Such results support the importance of data rather than methodology, and of quality over quantity. Who can assess the quality of this data? Data scientists provide expertise in analytics, but they do not have any experience with medical practice. The role should therefore be filled by scientists working in the medical field.
INTRODUCTION
- Whether we are considering EMR data or claim data, such as data generated by the National Health Insurance Service [1718] or Health Insurance Review & Assessment Service [1920], it is important to note that these are not data for research purposes [4]. Would it then be possible to conduct clinical research with data not intended for clinical research purposes? In order to secure good quality medical data, it is necessary to develop an operational definition and a strategy for DQM. If these are not properly considered, the reliability of the data will drop dramatically, and the results of research relying on this data will not be worth looking at.
- Conceptual definition vs. operational definition
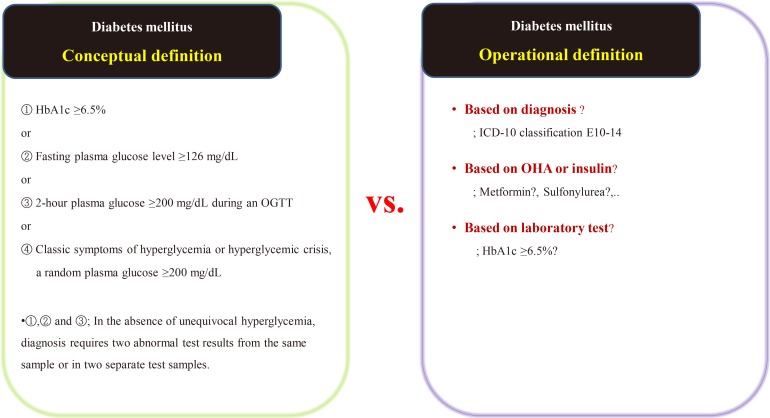
- Suppose we are conducting a study on the accompanying rate of cardiovascular disease in patients with diabetes mellitus (DM). The first thing to do is to define people with DM. The widely used definition of DM involves the patient's fasting/postprandial glucose and hemoglobin A1c (HbA1c) levels [2122]. This is a conceptual definition of DM, and most medical staff are familiar with it. In a data-driven study, however, this conceptual definition is not only impossible to extract, but also not very efficient. Therefore, we use operational rather than conceptual definition.
- We can develop an operational definition of DM using the International Classification of Diseases 10th Revision (ICD-10) classification code, an oral hypoglycemia agent such as metformin or sulfonylurea, or traditional laboratory findings such as HbA1c levels (Fig. 1) [23]. Of course, operational definitions can also be developed from the intersections or combinations produced by these three axes. There is no clear, correct way of developing an operational definition, and the researcher should do so based on their research objectives. It should be noted that, in real-world research, the first diagnosis date, the date of first taking the drug, and the date of blood test when diagnosed with DM are all different, as the data are often poorly stored. The study results are informed by the development of the operational definition, and this aspect of research requires the careful attention of medical staff. Importantly, steps need to be taken to minimize the bias that can occur in retrospective research using big data.
- Continuous communication between data scientists and medical staff can reduce research-related problems. However, medical staff often struggle to clarify an operational definition, and for this reason we need deeper reflection, rather than common sense, as the same data can render different results. Active collaboration with the medical staff who know the data best is required.
- Data quality management
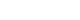
- After extracting medical data for analysis, it may become clear that it is not fit for research purposes. While data for research purposes must be expressed in a structured form, that is, in numbers, most EMR data is not well organized (Fig. 2). For example, result of hepatitis B surface antigen is described in various ways such as neg, negative, 0, and (−). In some cases, height and weight or systolic and diastolic blood pressure are reversed. Most clinicians will realize intuitively that the data in such cases were mistyped, but they are unable to change the numbers arbitrarily. Although some modifications can be made through operational definitions, in most cases such data is eventually converted to missing data. After all, DQM is the most time-consuming task in the research process, and must be managed by researcher or medical staff member who clearly understands medical data [4]. It is recommended that clear protocols and guidelines are put in place before the study so that other researchers can consistently follow the same approach.
HOW CAN WE ACQUIRE GOOD QUALITY DATA?
- As mentioned above, the advantages of big data are positively correlated with the amount of available data [9]. In the case of tertiary university hospitals, large amounts of meaningful medical data are expected to be available. However, for research conducted using real EMR data, the amount of missing data presents further challenges [2425]. Ultimately, the sample size from which the so-called big data can be extracted may contain smaller numbers than large-scaled randomized clinical trials. Given this, is the name “big data” still accurate, and how can greater amounts of medical data be collected? One answer would be to develop a multi-center registry as soon as possible to merge the available datasets from multiple hospitals [26].
- Big data research in a single institution presents various challenges, including controlling various operational definitions, DQM, and numerous biases [4]. While challenges exist with single hospitals, they are even larger when in a multi-center. Nevertheless, the integration of data from multiple hospitals appears to be required for future research. One critical consideration for the development of a multi-center registry is the development of an international coding system and mapping this [27], which would enable not only domestic partnerships but also international collaboration and would ultimately lead to the creation of an international standard [28]. Such a standard would include guidelines for the interpretation of specific types of data, and its establishment should be negotiated among various stakeholders, under the auspices of a suitably qualified institution. By doing this, every participating institution would agree to abide by the standard. However, in research conducted using multi-center registries, researchers tend not to consider the future need for an international code and choose instead a domestic standard that is agreed upon by a small number of institutions, rendering international mapping impossible. Thus, international mapping is mandatory if we are to consider future work.
- After the standard has been established, the role of hospitals and doctors need to be clarified. Inputting EMR data must be done according to the standardized format. Proper collection of data in the initial stages significantly decreases the time and money necessary for future research [4]. Data that is made or input without a specific standard will eventually lose its reliability and value. However, in the reality of the Korean medical environment, each patient is limited to 3 to 5 minutes when consulting with a doctor [29], during which it is realistically impossible for medical professionals to input data in the correct format. Nonetheless, the development of a standard remains important [28]; if EMR data accumulates in a standardized format, it will likely remain useful over time.
STANDARDIZATION AND THE DEVELOPMENT OF A MULTI-CENTER REGISTRY
- There has been increasing interest in CDM for research in various institutions in Korea. CDM unifies data in diverse formats into one common format [303132]. Among the various types of CDM, the one that is most popular in Korea is the observational health data sciences and informatics–observational medical outcomes partnership model (OHDSI-OMOP CDM), which is used in various institutions for retrospective big data collection and analysis, which makes it very useful for researchers [3133]. Moreover, CDM does not share patient identification numbers, which is a powerful advantage of CDM in terms of privacy However, the CDM itself has not been accredited by an authorized institution. Thus, there is a need to understand whether the CDM is an organizational or institutional standard, which is why the CDM consortium promises to use only in this consortium. Another challenge of CDM is that data mapping must be done manually, step-by-step [34]. Data mapping is nearly impossible to automatize as each hospital has a unique data structure, and automatic data mapping without consideration of the discrepancies would result in definite biases. EMR data does not exist solely for academic research, but is fundamentally aimed at supporting patient treatment [35]—a fact that should not be forgotten.
HOT ISSUES IN KOREA: THE COMMON DATA MODEL
- From the perspective of medical professionals, the purpose of collecting and standardizing medical big data needs to be defined clearly. Collecting standardized data properly is the most important task, as the main goal of retrospective research using EMR data is to care for patients. As long as data is collected properly, it does not matter whether you use deep learning, machine learning, or very simple statistical methods. Medical staff are best placed to use and interpret medical data, and the key is for them to critically observe the data, identifying any underlying problems and dealing with them appropriately. Finally, results comprising the most compelling value is found when refined data is combined with medical ideas.
CONCLUSIONS
-
Acknowledgements
- This work was supported by the Technology development Program (S2726209) funded by the Ministry of SMEs and Startups (MSS, Korea). I am indebted Prof Shin Soo-yong, whose lecture is referenced often in this text.
ACKNOWLEDGMENTS
-
CONFLICTS OF INTEREST: No potential conflict of interest relevant to this article was reported.
Article information
- 1. Hamet P, Tremblay J. Artificial intelligence in medicine. Metabolism 2017;69S:S36–S40. ArticlePubMed
- 2. Miller DD, Brown EW. Artificial intelligence in medical practice: the question to the answer? Am J Med 2018;131:129–133. ArticlePubMed
- 3. Kantarjian H, Yu PP. Artificial intelligence, big data, and cancer. JAMA Oncol 2015;1:573–574. ArticlePubMed
- 4. Kim HS, Kim JH. Proceed with caution when using real world data and real world evidence. J Korean Med Sci 2019;34:e28. ArticlePubMedPMC
- 5. Kim HS, Lee S, Kim JH. Real-world evidence versus randomized controlled trial: clinical research based on electronic medical records. J Korean Med Sci 2018;33:e213. ArticlePubMedPMC
- 6. Obermeyer Z, Emanuel EJ. Predicting the future: big data, machine learning, and clinical medicine. N Engl J Med 2016;375:1216–1219. ArticlePubMedPMC
- 7. Miotto R, Wang F, Wang S, Jiang X, Dudley JT. Deep learning for healthcare: review, opportunities and challenges. Brief Bioinform 2018;19:1236–1246. ArticlePubMedPDF
- 8. Ravi D, Wong C, Deligianni F, Berthelot M, Andreu-Perez J, Lo B, et al. Deep learning for health informatics. IEEE J Biomed Health Inform 2017;21:4–21. ArticlePubMed
- 9. Sun C, Shrivastava A, Singh S, Gupta A. Revisiting unreasonable effectiveness of data in deep learning era In: 2017 IEEE International Conference on Computer Vision (ICCV); 2017 Oct 22–29; Venice, IT. Piscataway, NJ: IEEE; 1997;pp 843–852.
- 10. Rubin DB. Estimating causal effects from large data sets using propensity scores. Ann Intern Med 1997;127:757–763. ArticlePubMed
- 11. Rajkomar A, Oren E, Chen K, Dai AM, Hajaj N, Hardt M, et al. Scalable and accurate deep learning with electronic health records. NPJ Digit Med 2018;1:18ArticlePubMedPMCPDF
- 12. Deo RC. Machine learning in medicine. Circulation 2015;132:1920–1930. ArticlePubMedPMC
- 13. Bastanlar Y, Ozuysal M. Introduction to machine learning. Methods Mol Biol 2014;1107:105–128. ArticlePubMed
- 14. Williams H. Microsoft's teen chatbot has gone wild [Internet]; Surry Hills: Gizmodo; 2016. cited 2019 Dec 9. Available from: https://www.gizmodo.com.au/2016/03/microsofts-teen-chatbot-has-gone-wild.
- 15. Levesque LE, Hanley JA, Kezouh A, Suissa S. Problem of immortal time bias in cohort studies: example using statins for preventing progression of diabetes. BMJ 2010;340:b5087. ArticlePubMed
- 16. Maugis PG. Big data uncertainties. J Forensic Leg Med 2018;57:7–11. ArticlePubMed
- 17. Lee J, Lee JS, Park SH, Shin SA, Kim K. Cohort profile: the National Health Insurance Service-National Sample Cohort (NHIS-NSC), South Korea. Int J Epidemiol 2017;46:e15. ArticlePubMedPDF
- 18. Noh J. The diabetes epidemic in Korea. Endocrinol Metab (Seoul) 2016;31:349–353. ArticlePubMedPMC
- 19. Seo GH, Chung JH. Incidence and prevalence of overt hypothyroidism and causative diseases in Korea as determined using claims data provided by the Health Insurance Review and Assessment Service. Endocrinol Metab (Seoul) 2015;30:288–296. ArticlePubMedPMC
- 20. Lee YK, Yoon BH, Koo KH. Epidemiology of osteoporosis and osteoporotic fractures in South Korea. Endocrinol Metab (Seoul) 2013;28:90–93. ArticlePubMedPMC
- 21. American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 2004;27:S5–S10. ArticlePubMed
- 22. Ko SH, Hur KY, Rhee SY, Kim NH, Moon MK, Park SO, et al. Antihyperglycemic agent therapy for adult patients with type 2 diabetes mellitus 2017: a position statement of the Korean Diabetes Association. Diabetes Metab J 2017;41:337–348. ArticlePubMedPMC
- 23. International Expert Committee. International Expert Committee report on the role of the A1C assay in the diagnosis of diabetes. Diabetes Care 2009;32:1327–1334. ArticlePubMedPMC
- 24. Zhang Z. Missing data exploration: highlighting graphical presentation of missing pattern. Ann Transl Med 2015;3:356PubMedPMC
- 25. Kim TM, Kim H, Jeong YJ, Baik SJ, Yang SJ, Lee SH, et al. The differences in the incidence of diabetes mellitus and prediabetes according to the type of HMG-CoA reductase inhibitors prescribed in Korean patients. Pharmacoepidemiol Drug Saf 2017;26:1156–1163. ArticlePubMed
- 26. Chen PH, Loehfelm TW, Kamer AP, Lemmon AB, Cook TS, Kohli MD. Toward data-driven radiology education-early experience building multi-institutional academic trainee interpretation log database (MATILDA). J Digit Imaging 2016;29:638–644. ArticlePubMedPMCPDF
- 27. Matney SA, Settergren TT, Carrington JM, Richesson RL, Sheide A, Westra BL. Standardizing physiologic assessment data to enable big data analytics. West J Nurs Res 2017;39:63–77. ArticlePubMed
- 28. Kalra D. Electronic health record standards. Yearb Med Inform 2006;136–144. ArticlePubMedPDF
- 29. Lee CH, Lim H, Kim Y, Park AH, Park EC, Kang JG. Analysis of appropriate outpatient consultation time for clinical departments. Health Policy Manag 2014;24:254–260.ArticlePDF
- 30. Kim H, Choi J, Jang I, Quach J, Ohno-Machado L. Feasibility of representing data from published nursing research using the OMOP common data model. AMIA Annu Symp Proc 2017;2016:715–723. PubMedPMC
- 31. Ceusters W, Blaisure J. A realism-based view on counts in OMOP’s common data model. Stud Health Technol Inform 2017;237:55–62. PubMed
- 32. Kimura E, Suzuki H. Development of a common data model facilitating clinical decision-making and analyses. Stud Health Technol Inform 2019;264:1514–1515. PubMed
- 33. Park RW. The distributed research network, observational health data sciences and informatics, and the South Korean research network. Korean J Med 2019;94:309–314.ArticlePDF
- 34. FitzHenry F, Resnic FS, Robbins SL, Denton J, Nookala L, Meeker D, et al. Creating a common data model for comparative effectiveness with the observational medical outcomes partnership. Appl Clin Inform 2015;6:536–547. ArticlePubMedPMCPDF
- 35. Kim HS. Decision-making in artificial intelligence: is it always correct? J Korean Med Sci 2020 In Press.Article
References
Example of the conceptual [23] and operational definitions of diabetes mellitus. HbA1c, hemoglobin A1c; OGTT, oral glucose tolerance test; ICD-10, International Classification of Diseases 10th Revision; OHA, oral hypoglycemic agents.
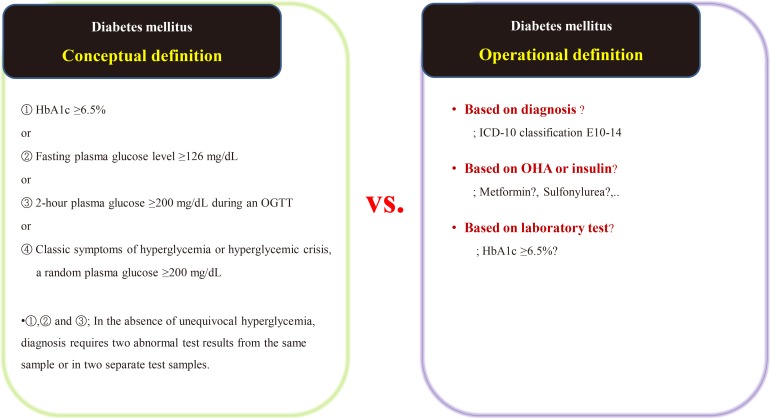

Examples of real cases requiring data quality management. (A) Various written data. Even though the laboratory test result is “<3,” it is also written as “1.” The physicians' role is defining and classifying data, and staff who are most familiar with the data should do so. (B) Example of incorrectly entered data. AST, aspartate aminotransferase; GOT, glutamate oxaloacetate transaminase; ALT, aspartate aminotransferase; GPT, glutamate pyruvate transaminase; HBsAg, hepatitis B surface antigen; HBsAb, hepatitis B surface antibody; HDL, high-density lipoprotein; LDL, low-density lipoprotein.
Figure & Data
References
Citations

- Current status of remote collaborative care for hypertension in medically underserved areas
Seo Yeon Baik, Kyoung Min Kim, Hakyoung Park, Jiwon Shinn, Hun-Sung Kim
Cardiovascular Prevention and Pharmacotherapy.2024; 6(1): 33. CrossRef - Prediction of Cardiovascular Complication in Patients with Newly Diagnosed Type 2 Diabetes Using an XGBoost/GRU-ODE-Bayes-Based Machine-Learning Algorithm
Joonyub Lee, Yera Choi, Taehoon Ko, Kanghyuck Lee, Juyoung Shin, Hun-Sung Kim
Endocrinology and Metabolism.2024; 39(1): 176. CrossRef - Dark Data in Real-World Evidence: Challenges, Implications, and the Imperative of Data Literacy in Medical Research
Hun-Sung Kim
Journal of Korean Medical Science.2024;[Epub] CrossRef - A comparative analysis: health data protection laws in Malaysia, Saudi Arabia and EU General Data Protection Regulation (GDPR)
Jawahitha Sarabdeen, Mohamed Mazahir Mohamed Ishak
International Journal of Law and Management.2024;[Epub] CrossRef - Long-Term Risk of Cardiovascular Disease Among Type 2 Diabetes Patients According to Average and Visit-to-Visit Variations of HbA1c Levels During the First 3 Years of Diabetes Diagnosis
Hyunah Kim, Da Young Jung, Seung-Hwan Lee, Jae-Hyoung Cho, Hyeon Woo Yim, Hun-Sung Kim
Journal of Korean Medical Science.2023;[Epub] CrossRef - Comparison of cardiocerebrovascular disease incidence between angiotensin converting enzyme inhibitor and angiotensin receptor blocker users in a real-world cohort
Suehyun Lee, Hyunah Kim, Hyeon Woo Yim, Kim Hun-Sung, Ju Han Kim
Journal of Applied Biomedicine.2023; 21(1): 7. CrossRef - Multi-Omics and Management of Follicular Carcinoma of the Thyroid
Thifhelimbilu Emmanuel Luvhengo, Ifongo Bombil, Arian Mokhtari, Maeyane Stephens Moeng, Demetra Demetriou, Claire Sanders, Zodwa Dlamini
Biomedicines.2023; 11(4): 1217. CrossRef - Correlation analysis of cancer incidence after pravastatin treatment
Jin Yu, Raeun Kim, Jiwon Shinn, Man Young Park, Hun-Sung Kim
Cardiovascular Prevention and Pharmacotherapy.2023; 5(2): 61. CrossRef - A New Strategy for Evaluating the Quality of Laboratory Results for Big Data Research: Using External Quality Assessment Survey Data (2010–2020)
Eun-Jung Cho, Tae-Dong Jeong, Sollip Kim, Hyung-Doo Park, Yeo-Min Yun, Sail Chun, Won-Ki Min
Annals of Laboratory Medicine.2023; 43(5): 425. CrossRef - Weight loss and side-effects of liraglutide and lixisenatide in obesity and type 2 diabetes mellitus
Jeongmin Lee, Raeun Kim, Min-Hee Kim, Seung-Hwan Lee, Jae-Hyoung Cho, Jung Min Lee, Sang-Ah Jang, Hun-Sung Kim
Primary Care Diabetes.2023; 17(5): 460. CrossRef - Cohort profile for development of machine learning models to predict healthcare-related adverse events (Demeter): clinical objectives, data requirements for modelling and overview of data set for 2016–2018
Svetlana Artemova, Ursula von Schenck, Rui Fa, Daniel Stoessel, Hadiseh Nowparast Rostami, Pierre-Ephrem Madiot, Jean-Marie Januel, Daniel Pagonis, Caroline Landelle, Meghann Gallouche, Christophe Cancé, Frederic Olive, Alexandre Moreau-Gaudry, Sigurd Pri
BMJ Open.2023; 13(8): e070929. CrossRef - The Present and Future of Artificial Intelligence-Based Medical Image in Diabetes Mellitus: Focus on Analytical Methods and Limitations of Clinical Use
Ji-Won Chun, Hun-Sung Kim
Journal of Korean Medical Science.2023;[Epub] CrossRef - Construction and application on the training course of information literacy for clinical nurses
Chao Wu, Yinjuan Zhang, Jing Wu, Linyuan Zhang, Juan Du, Lu Li, Nana Chen, Liping Zhu, Sheng Zhao, Hongjuan Lang
BMC Medical Education.2023;[Epub] CrossRef - Lightweight Histological Tumor Classification Using a Joint Sparsity-Quantization Aware Training Framework
Dina Aboutahoun, Rami Zewail, Keiji Kimura, Mostafa I. Soliman
IEEE Access.2023; 11: 119342. CrossRef - Long-Term Cumulative Exposure to High γ-Glutamyl Transferase Levels and the Risk of Cardiovascular Disease: A Nationwide Population-Based Cohort Study
Han-Sang Baek, Bongseong Kim, Seung-Hwan Lee, Dong-Jun Lim, Hyuk-Sang Kwon, Sang-Ah Chang, Kyungdo Han, Jae-Seung Yun
Endocrinology and Metabolism.2023; 38(6): 770. CrossRef - Comorbidity Patterns and Management in Inpatients with Endocrine Diseases by Age Groups in South Korea: Nationwide Data
Sung-Soo Kim, Hun-Sung Kim
Journal of Personalized Medicine.2023; 14(1): 42. CrossRef - Angiotensin‐converting enzyme inhibitors versus angiotensin receptor blockers: New‐onset diabetes mellitus stratified by statin use
Juyoung Shin, Hyunah Kim, Hyeon Woo Yim, Ju Han Kim, Suehyun Lee, Hun‐Sung Kim
Journal of Clinical Pharmacy and Therapeutics.2022; 47(1): 97. CrossRef - Physician Knowledge Base: Clinical Decision Support Systems
Sira Kim, Eung-Hee Kim, Hun-Sung Kim
Yonsei Medical Journal.2022; 63(1): 8. CrossRef - Sodium-Glucose Cotransporter-2 Inhibitor-Related Diabetic Ketoacidosis: Accuracy Verification of Operational Definition
Dong Yoon Kang, Hyunah Kim, SooJeong Ko, HyungMin Kim, Jiwon Shinn, Min-Gyu Kang, Sun-ju Byeon, Jeong-Hee Choi, Soo-Yong Shin, Hun-Sung Kim
Journal of Korean Medical Science.2022;[Epub] CrossRef - Drug Repositioning: Exploring New Indications for Existing Drug-Disease Relationships
Hun-Sung Kim
Endocrinology and Metabolism.2022; 37(1): 62. CrossRef - A Study on Methodologies of Drug Repositioning Using Biomedical Big Data: A Focus on Diabetes Mellitus
Suehyun Lee, Seongwoo Jeon, Hun-Sung Kim
Endocrinology and Metabolism.2022; 37(2): 195. CrossRef - Development of a predictive model for the side effects of liraglutide
Jiyoung Min, Jiwon Shinn, Hun-Sung Kim
Cardiovascular Prevention and Pharmacotherapy.2022; 4(2): 87. CrossRef - Understanding and Utilizing Claim Data from the Korean National Health Insurance Service (NHIS) and Health Insurance Review & Assessment (HIRA) Database for Research
Dae-Sung Kyoung, Hun-Sung Kim
Journal of Lipid and Atherosclerosis.2022; 11(2): 103. CrossRef - The Impact of the Association between Cancer and Diabetes Mellitus on Mortality
Sung-Soo Kim, Hun-Sung Kim
Journal of Personalized Medicine.2022; 12(7): 1099. CrossRef - Development of Various Diabetes Prediction Models Using Machine Learning Techniques
Juyoung Shin, Jaewon Kim, Chanjung Lee, Joon Young Yoon, Seyeon Kim, Seungjae Song, Hun-Sung Kim
Diabetes & Metabolism Journal.2022; 46(4): 650. CrossRef - Characteristics of Glycemic Control and Long-Term Complications in Patients with Young-Onset Type 2 Diabetes
Han-sang Baek, Ji-Yeon Park, Jin Yu, Joonyub Lee, Yeoree Yang, Jeonghoon Ha, Seung Hwan Lee, Jae Hyoung Cho, Dong-Jun Lim, Hun-Sung Kim
Endocrinology and Metabolism.2022; 37(4): 641. CrossRef - Retrospective cohort analysis comparing changes in blood glucose level and body composition according to changes in thyroid‐stimulating hormone level
Hyunah Kim, Da Young Jung, Seung‐Hwan Lee, Jae‐Hyoung Cho, Hyeon Woo Yim, Hun‐Sung Kim
Journal of Diabetes.2022; 14(9): 620. CrossRef - Long-Term Changes in HbA1c According to Blood Glucose Control Status During the First 3 Months After Visiting a Tertiary University Hospital
Hyunah Kim, Da Young Jung, Seung-Hwan Lee, Jae-Hyoung Cho, Hyeon Woo Yim, Hun-Sung Kim
Journal of Korean Medical Science.2022;[Epub] CrossRef - Medication based machine learning to identify subpopulations of pediatric hemodialysis patients in an electronic health record database
Autumn M. McKnite, Kathleen M. Job, Raoul Nelson, Catherine M.T. Sherwin, Kevin M. Watt, Simon C. Brewer
Informatics in Medicine Unlocked.2022; 34: 101104. CrossRef - Improving Machine Learning Diabetes Prediction Models for the Utmost Clinical Effectiveness
Juyoung Shin, Joonyub Lee, Taehoon Ko, Kanghyuck Lee, Yera Choi, Hun-Sung Kim
Journal of Personalized Medicine.2022; 12(11): 1899. CrossRef - A Study on Weight Loss Cause as per the Side Effect of Liraglutide
Jin Yu, Jeongmin Lee, Seung-Hwan Lee, Jae-Hyung Cho, Hun-Sung Kim, Heng Zhou
Cardiovascular Therapeutics.2022; 2022: 1. CrossRef - Risk Classification and Subphenotyping of Acute Kidney Injury: Concepts and Methodologies
Javier A. Neyra, Jin Chen, Sean M. Bagshaw, Jay L. Koyner
Seminars in Nephrology.2022; 42(3): 151285. CrossRef - Estimation of sodium‐glucose cotransporter 2 inhibitor–related genital and urinary tract infections via electronic medical record–based common data model
SooJeong Ko, HyungMin Kim, Jiwon Shinn, Sun‐ju Byeon, Jeong‐Hee Choi, Hun‐Sung Kim
Journal of Clinical Pharmacy and Therapeutics.2021; 46(4): 975. CrossRef - Blood glucose levels and bodyweight change after dapagliflozin administration
Hyunah Kim, Seung‐Hwan Lee, Hyunyong Lee, Hyeon Woo Yim, Jae‐Hyoung Cho, Kun‐Ho Yoon, Hun‐Sung Kim
Journal of Diabetes Investigation.2021; 12(9): 1594. CrossRef - Artificial intelligence in healthcare: possibilities of patent protection
T. N. Erivantseva, Yu. V. Blokhina
FARMAKOEKONOMIKA. Modern Pharmacoeconomic and Pharmacoepidemiology.2021; 14(2): 270. CrossRef - Lack of Acceptance of Digital Healthcare in the Medical Market: Addressing Old Problems Raised by Various Clinical Professionals and Developing Possible Solutions
Jong Il Park, Hwa Young Lee, Hyunah Kim, Jisan Lee, Jiwon Shinn, Hun-Sung Kim
Journal of Korean Medical Science.2021;[Epub] CrossRef - Prospect of Artificial Intelligence Based on Electronic Medical Records
Suehyun Lee, Hun-Sung Kim
Journal of Lipid and Atherosclerosis.2021; 10(3): 282. CrossRef - Data Pseudonymization in a Range That Does Not Affect Data Quality: Correlation with the Degree of Participation of Clinicians
Soo-Yong Shin, Hun-Sung Kim
Journal of Korean Medical Science.2021;[Epub] CrossRef - Development of a Predictive Model for Glycated Hemoglobin Values and Analysis of the Factors Affecting It
HyeongKyu Park, Da Young Lee, So young Park, Jiyoung Min, Jiwon Shinn, Dae Ho Lee, Soon Hyo Kwon, Hun-Sung Kim, Nan Hee Kim
Cardiovascular Prevention and Pharmacotherapy.2021; 3(4): 106. CrossRef - Modeling of Changes in Creatine Kinase after HMG-CoA Reductase Inhibitor Prescription
Hun-Sung Kim, Jiyoung Min, Jiwon Shinn, Oak-Kee Hong, Jang-Won Son, Seong-Su Lee, Sung-Rae Kim, Soon Jib Yoo
Cardiovascular Prevention and Pharmacotherapy.2021; 3(4): 115. CrossRef - TRAINING IN BIG DATA TECHNOLOGIES OF MEDICAL UNIVERSITY STUDENTS
K.S ITINSON
AZIMUTH OF SCIENTIFIC RESEARCH: PEDAGOGY AND PSYCHOLOGY.2021;[Epub] CrossRef - Machine Learning Applications in Endocrinology and Metabolism Research: An Overview
Namki Hong, Heajeong Park, Yumie Rhee
Endocrinology and Metabolism.2020; 35(1): 71. CrossRef - Lessons from Use of Continuous Glucose Monitoring Systems in Digital Healthcare
Hun-Sung Kim, Kun-Ho Yoon
Endocrinology and Metabolism.2020; 35(3): 541. CrossRef - Apprehensions about Excessive Belief in Digital Therapeutics: Points of Concern Excluding Merits
Hun-Sung Kim
Journal of Korean Medical Science.2020;[Epub] CrossRef - Medical Ethics in the Era of Artificial Intelligence Based on Medical Big Data
Hae-Ran Na, Hun-Sung Kim
The Journal of Korean Diabetes.2020; 21(3): 126. CrossRef - Machine Learning Application in Diabetes and Endocrine Disorders
Namki Hong, Heajeong Park, Yumie Rhee
The Journal of Korean Diabetes.2020; 21(3): 130. CrossRef - Real World Data and Artificial Intelligence in Diabetology
Kwang Joon Kim
The Journal of Korean Diabetes.2020; 21(3): 140. CrossRef

 KES
KES
 PubReader
PubReader ePub Link
ePub Link Cite
Cite