The Presence of Clonal Hematopoiesis Is Negatively Associated with Diabetic Peripheral Neuropathy in Type 2 Diabetes
Article information
Abstract
Background
Clonal hematopoiesis of indeterminate potential (CHIP) has been reported to be associated with increased cardiovascular disease, aging and insulin resistance. Despite the debate of causal contribution of CHIP on metabolic diseases, we want to explore whether CHIP is related to diabetic peripheral neuropathy (DPN).
Methods
This study analyzed the prevalence of CHIP in patients with type 2 diabetes classified according to DPN status. Logistic regression analysis was used to evaluate the association between CHIP and DPN.
Results
CHIP was more prevalent in subjects without DPN than those with DPN (19.9% vs. 8.8%, respectively; P=0.013). Individuals having any CHIP, or DNA methyltransferase 3A (DNMT3A) CHIP were less likely to have any abnormality shown in DPN test; the adjusted odds ratio were 0.85 (95% confidence interval [CI], 0.73 to 1.00) and 0.70 (95% CI, 0.56 to 0.89), respectively. Interestingly, DNMT3A CHIP showed the negative association, but Tet methylcytosine dioxygenase 2 (TET2) CHIP showed the positive association with abnormal feet electrochemical skin conductance level.
Conclusion
On the contrary to expectations, CHIP was negatively associated with DPN. Functional linking between the mutation in hematopoietic cells and DPN, and the opposite role of DNMT3A and TET2 should be investigated.
INTRODUCTION
Clonal hematopoiesis of indeterminate potential (CHIP) is defined as an expansion of hematopoietic stem cells (HSCs) driven by somatic mutations without the development of any hematologic malignancy [1]. CHIP has been reported to be associated with aging or stress to the bone marrow, including that caused by radiation or chemotherapy [2], and to lead to in increased all-cause mortality, possibly because of a higher risk of hematologic malignancy or cardiovascular disease [3]. Somatic chromosomal alterations in peripheral blood cells are also associated with an increased risk of type 2 diabetes, even after adjustment for confounding factors [3]. Increased insulin resistance, altered immune function, and increased inflammation may be underlying mechanisms related to the poor health outcomes of CHIP [1].
Diabetic peripheral neuropathy (DPN) shares common risk factors with cardiovascular disease in people with type 1 diabetes [4], and this association has been observed in Koreans with type 2 diabetes [5]. From the mechanistic view, endothelial dysfunction, an early vascular abnormality, was observed in both DPN and cardiovascular disease [6]. We hypothesized that CHIP might be more prevalent in subjects with DPN compared to their counterpart. However, whether CHIP is related to DPN, an important microvascular complication of diabetes, remains unclear. In this study, we investigated the prevalence of CHIP in subjects with type 2 diabetes classified according to DPN status and analyzed the associations between CHIP and DPN.
METHODS
Study population
We enrolled adult subjects with type 2 diabetes from a tertiary academic hospital since November 2016 [7]. The original prospective observational study was designed to develop reliable tools and biomarkers for identifying DPN. We annually recruited 100 subjects and this study used the data from the third wave survey. Among them we collected DNA from 294 subjects. This study was approved by the Institutional Review Board of Seoul National University Bundang Hospital (IRB No.: B-2106/688-303). Informed consent was waived by the board.
Assessment of DPN
DPN was evaluated using the Michigan Neuropathy Screening Instrument-questionnaire (MNSI-Q) and -physical examination (MNSI-PE), a 10-g monofilament test (Semmes-Weinstein 5.07 monofilaments), and measurement of feet electrochemical skin conductance (ESC) (SUDOSCAN, Impeto Medical, Paris, France). A research nurse performed aforementioned evaluations with standard operation procedure. The cut-off values of MNSI-Q and MNSI-PE were adopted from the original study which firstly developed MSNI [7]. We used cut-off values of a 10-g monofilament test and ESC according to the study from Korean population [8,9].
CHIP analysis
Genomic DNA was separated from peripheral blood. Targeted sequencing was performed using a custom panel consisting of 89 genes frequently involved in CHIP, including DNA methyltransferase 3A (DNMT3A), Tet methylcytosine dioxygenase 2 (TET2), ASXL transcription regulator (ASXL1), Janus kinase 2 (JAK2), and tumor protein p53 (TP53) [10] with an average target-depth of 1,100×. All nonsynonymous variants with a variant allele frequency of 2% to 30% were considered to be CHIP variants. Common germline variants listed in the gnomAD, 100-0 Genomes v3, ESP6500, and ExAC databases, and an internal panel of 1,000 Koreans were excluded.
Statistical analysis
The chi-square test was used to compare the prevalence of CHIP between subjects with and without DPN. Odds ratios (ORs) and 95% confidence interval (CIs) were estimated using univariable and multivariable logistic regression analysis. Covariates were selected based on univariable logistic analysis showing a significant association with DPN. Statistical analysis was performed using IBM SPSS version 25.0 (IBM Corp., Armonk, NY, USA) and Python version 3.7.9 (Python Software Foundation, Wilmington, DE, USA).
RESULTS
Clinical and biochemical characteristics
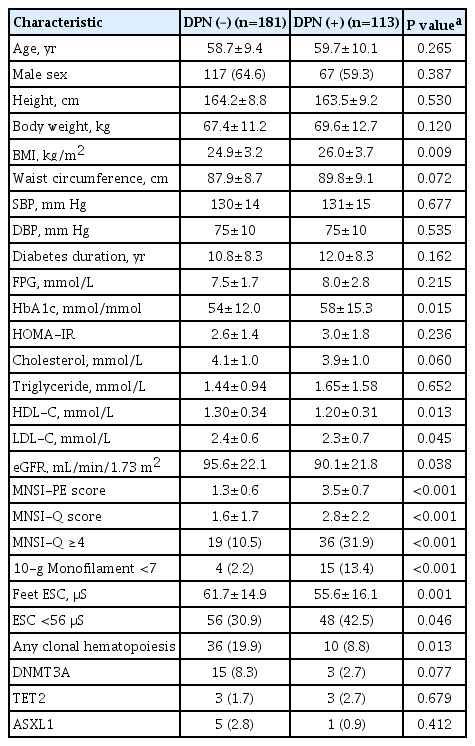
Among the 294 subjects (184 men; mean age 59.1 years), 113 (38.4%) had an MNSI-PE score >2.0, and 183 (67.3%) had any abnormality identified in DPN tests. Compared with subjects without DPN, those with DPN (MNSI-PE >2.0) were more obese, had higher hemoglobin A1c (HbA1c) and lower high-density lipoprotein cholesterol (HDL-C) levels, as well as a lower estimated glomerular filtration rate (eGFR) (Table 1).
Parameters associated with DPN
According to univariable logistic regression analysis, body mass index (OR, 1.098; 95% CI, 1.023 to 1.179), and HbA1c (OR, 1.264; 95% CI, 1.045 to 1.529) were positively associated with the presence of DPN (Table 2). In contrast, HDL-C (OR, 0.975; 95% CI, 0.955 to 0.997), and eGFR (OR, 0.989; 95% CI, 0.978 to 0.999) showed negative association with DPN.
The prevalence of CHIP and its association with DPN
CHIP was more frequently detected in subjects without DPN than those with DPN (19.9% vs. 8.8%, respectively; P=0.013). The most prevalent variants of CHIP occurred in DNMT3A and TET2 (Table 1). After multivariable adjustment for body mass index, HbA1c, HDL-C and eGFR, CHIP and DNMT3A CHIP were independently associated with any abnormality in DPN test; the adjusted ORs were 0.85 (95% CI, 0.73 to 1.00) and 0.70 (95% CI, 0.56 to 0.89), respectively (Fig. 1). However, there was no statistical significance between TET2 CHIP and any abnormal DPN. Interestingly, subjects with DNMT3A and TET2 CHIP showed the opposite association with abnormal feet ESC level; adjusted OR of DNMT3A and TET2 were 0.74 (95% CI, 0.59 to 093) and 1.57 (95% CI, 1.07 to 2.32), respectively.

Adjusted odds ratio (OR) (95% confidence interval [CI]) for the associations between clonal hematopoiesis and the presence of abnormalities in diabetic peripheral neuropathy (DPN). (A) All clonal hematopoiesis, (B) DNMT3A clonal hematopoiesis, and (C) TET2 clonal hematopoiesis. The OR was adjusted for body mass index, hemoglobin A1c, and high-density lipoprotein cholesterol levels, and estimated glomerular filtration rate. CH, clonal hematopoiesis; MNSI-Q, Michigan Neuropathy Screening Instrument-questionnaire; MNSI-PE, Michigan Neuropathy Screening Instrument-physical examination; MF, monofilament test; ESC, electrochemical skin conductance; DNMT3A, DNA methyltransferase 3A; TET2, Tet methylcytosine dioxygenase 2.
DISCUSSION
To our knowledge, this is the first report on the association between CHIP and DPN in subjects with type 2 diabetes. Our findings suggest a possible negative association between CHIP and DPN. This was an unexpected finding because previous observational studies have reported that CHIP is associated with poor health outcomes for cardiometabolic diseases [3,11]. Further mechanistic studies are needed to identify the causes and consequences of CHIP in DPN and to investigate the true role of CHIP in DPN.
DNMT3A and TET2 are more frequently observed in association with CHIP, based on the results from our study and others [3,12]. These two genes encode for enzymes with opposing actions in DNA methylation. DNMT3A is responsible for de novo methylation of cytosine bases in DNA, whereas TET2 is involved in DNA demethylation [13]. At the cellular level, although both TET2 and DNMT3A loss-of-function mutations provide a growth advantage in HSCs, these genes have different effects on HSC differentiation. The TET2 mutation promotes HSC myeloid differentiation, whereas the DNMT3A mutation inhibits differentiation of HSCs [14]. In addition, the TET2 loss-of-function mutation in hematopoietic cells induces systemic insulin resistance in aged and obese mice [15]. However, the role of the DNMT3A mutation in HSCs in systemic insulin resistance or inflammation has not been confirmed.
Peripheral nerves have the capacity for regeneration, and epigenetic mechanisms are involved in transcriptional reprogramming [16]. We could not suggest the exact mechanism of CHIP in the development of DPN from our findings, but we have to consider the influence of counterbalance, between DNA methylation and demethylation, on DPN. The critical targets of epigenetic modification must be identified to understand the detailed mechanisms responsible for the epigenetic changes in DPN. For example, a previous study reported that RG108, a DNMT inhibitor, attenuated thermal hyperalgesia via upregulation of the mu opioid receptor in a mouse model of neuropathic pain [17]. By contrast, TET3 knockout in neural progenitor cells has been shown to produce rapid neural apoptosis and to impair the terminal differentiation of neurons [18]. Epigenetic changes in neurons may have various pathogenic mechanisms depending on the type of gene or the stage of differentiation. Further study to discover the contrasting action of DNMT and TET3 in DPN progression will be necessary.
SUDOSCAN can detect abnormality of unmyelinated c-fiber function which innervates sweat gland [19]. Selvarajah et al. [19] reported that SUDOSCAN can classify DPN and CAN with sensitivity of 87.5% and 76.2% of specificity, and with sensitivity of 65.0% and specificity of 80.0%, respectively. If the patient had abnormal small-fiber neuropathy, we can detect the low ESC level. In our study, we identified that DNMT3A was associated with normal ESC levels; however, TET2 was associated with decrease of ESC levels. This opposite association was just shown in DPN test for SUDOSCAN. In this regards, CHIP might primarily involve in one of features of small-fiber neuropathy rather than large-fiber neuropathy. Therefore, we could develop a screening tool for DPN using CHIP analysis.
The limitation of our study include the small sample size, and given the exploratory nature of the current study, we did not calculate exact sample size. We could not identify consistent statistically significant associations between each CHIP and individual DPN test results. Further validation studies including more subjects are needed. In addition, our study lacked data relating to the phenotype of DPN (e.g., painless or painful), and we only performed limited tests for DPN [20]. For example, we just adopted SUDOSCAN for small-fiber neuropathy, and we did not perform other tools such as pinprick test and quantitative sensory test of temperature or pain. Finally, the cross-sectional nature of this study did not allow us to draw inferences about the causality of CHIP in DPN development. Despite these limitations, the present study provides the first human data suggesting possible interactions between CHIP and DPN.
In conclusion, the present study provides evidence of a generally protective role of CHIP in DPN and possible opposite effects of DMNT3A and TET2 CHIP in type 2 diabetes. Further mechanistic studies of the epigenetic modifications of the target genes may provide useful targets for DPN treatment.
Notes
CONFLICTS OF INTEREST
Youngil Koh and Han Song are cofounders of Genome Opinion Inc. and hold stock in the company.
AUTHOR CONTRIBUTIONS
Conception or design: T.J.O., Y.K., S.H.C. Acquisition, analysis, or interpretation of data: T.J.O., H.S. Drafting the work or revising: T.J.O., Y.K., S.H.C. Final approval of the manuscript: T.J.O., H.S., Y.K., S.H.C.
Acknowledgements
Genome opinion Inc. provided the targeted panel (LifeEx CH panel) sequencing and CHIP variant analysis.