Articles
- Page Path
- HOME > Endocrinol Metab > Volume 33(2); 2018 > Article
-
Review ArticleGenetic Polymorphism Predisposing to Differentiated Thyroid Cancer: A Review of Major Findings of the Genome-Wide Association Studies
-
Vladimir A. Saenko1
 , Tatiana I. Rogounovitch2
, Tatiana I. Rogounovitch2 -
Endocrinology and Metabolism 2018;33(2):164-174.
DOI: https://doi.org/10.3803/EnM.2018.33.2.164
Published online: June 21, 2018
1Department of Radiation Molecular Epidemiology, Atomic Bomb Disease Institute, Nagasaki University, Nagasaki, Japan.
2Department of Global Health, Medicine and Welfare, Atomic Bomb Disease Institute, Nagasaki University, Nagasaki, Japan.
- Corresponding author: Vladimir A. Saenko. Department of Radiation Molecular Epidemiology, Atomic Bomb Disease Institute, Nagasaki University, 1-12-4 Sakamoto, Nagasaki 852-8523, Japan. Tel: +81-95-819-7122, Fax: +81-95-819-7169, saenko@nagasaki-u.ac.jp
• Received: April 24, 2018 • Revised: May 4, 2018 • Accepted: May 10, 2018
Copyright © 2018 Korean Endocrine Society
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- ABSTRACT
- INTRODUCTION
- SINGLE-NUCLEOTIDE POLYMORPHISMS IN HUMAN GENOME AND THE GENETIC COMPONENT OF NON-SYNDROMIC THYROID CANCER
- TYPES AND METHODOLOGY OF GENETIC ASSOCIATION STUDIES
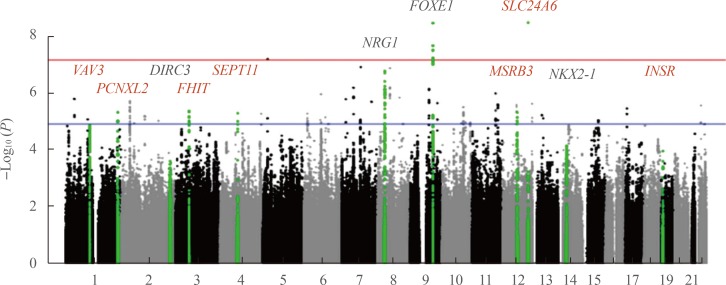
- MAJOR GWAS FINDINGS IN DIFFERENTIATED THYROID CANCER
- CUMULATIVE EFFECT AND PREDICTION ABILITY OF RISK-CONFERRING SNPs
- CONCLUSIONS
- ACKNOWLEDGMENTS
- Article information
- References
Figure & Data
References
Citations
Citations to this article as recorded by 

- Radiation-Related Thyroid Cancer
Vladimir Saenko, Norisato Mitsutake
Endocrine Reviews.2024; 45(1): 1. CrossRef - Gene variants polymorphisms and uterine leiomyoma: an updated review
Sonal Upadhyay, Pawan K. Dubey
Frontiers in Genetics.2024;[Epub] CrossRef - Disease-associated non-coding variants alter NKX2-5 DNA-binding affinity
Edwin G. Peña-Martínez, Alejandro Rivera-Madera, Diego A. Pomales-Matos, Leandro Sanabria-Alberto, Brittany M. Rosario-Cañuelas, Jessica M. Rodríguez-Ríos, Emanuel A. Carrasquillo-Dones, José A. Rodríguez-Martínez
Biochimica et Biophysica Acta (BBA) - Gene Regulatory Mechanisms.2023; 1866(1): 194906. CrossRef - The Role of Genetic Polymorphisms in Differentiated Thyroid Cancer: A 2023 Update
Robert Aurelian Tiucă, Oana Mirela Tiucă, Ionela Maria Pașcanu
Biomedicines.2023; 11(4): 1075. CrossRef - Genetic susceptibility to hereditary non-medullary thyroid cancer
Tina Kamani, Parsa Charkhchi, Afshan Zahedi, Mohammad R. Akbari
Hereditary Cancer in Clinical Practice.2022;[Epub] CrossRef - Genetic signature of differentiated thyroid carcinoma susceptibility: a machine learning approach
Giulia Brigante, Clara Lazzaretti, Elia Paradiso, Federico Nuzzo, Martina Sitti, Frank Tüttelmann, Gabriele Moretti, Roberto Silvestri, Federica Gemignani, Asta Försti, Kari Hemminki, Rossella Elisei, Cristina Romei, Eric Adriano Zizzi, Marco Agostino Der
European Thyroid Journal.2022;[Epub] CrossRef - Study of single nucleotide polymorphism of vascular endothelium factor in patients with differentiated thyroid cancer
Mohamad Mohsen Motawea, Maysaa El Sayed Zaki, Maha Saif, Asmaa Osama BS Osman, Aml Mohamed Nada
Clinical Diabetes and Endocrinology.2022;[Epub] CrossRef - The Contribution of Genetic Variants to the Risk of Papillary Thyroid Carcinoma in the Kazakh Population: Study of Common Single Nucleotide Polymorphisms and Their Clinicopathological Correlations
Zhanna Mussazhanova, Tatiana I. Rogounovitch, Vladimir A. Saenko, Ainur Krykpayeva, Maira Espenbetova, Bauyrzhan Azizov, Hisayoshi Kondo, Katsuya Matsuda, Zhanna Kalmatayeva, Raushan Issayeva, Zhanar Yeleubayeva, Madina Madiyeva, Aray Mukanova, Marat Sand
Frontiers in Endocrinology.2021;[Epub] CrossRef - Incidence of the CHEK2 Germline Mutation and Its Impact on Clinicopathological Features, Treatment Responses, and Disease Course in Patients with Papillary Thyroid Carcinoma
Danuta Gąsior-Perczak, Artur Kowalik, Krzysztof Gruszczyński, Agnieszka Walczyk, Monika Siołek, Iwona Pałyga, Sławomir Trepka, Estera Mikina, Tomasz Trybek, Janusz Kopczyński, Agnieszka Suligowska, Rafał Ślusarczyk, Agnieszka Gonet, Jarosław Jaskulski, Pa
Cancers.2021; 13(3): 470. CrossRef - Carcinoma diferenciado de tiroides familiar: más allá de las formas sindrómicas
Aida Orois, Mireia Mora, Irene Halperin, Josep Oriola
Endocrinología, Diabetes y Nutrición.2021; 68(4): 260. CrossRef - Gene network and biological pathways associated with susceptibility to differentiated thyroid carcinoma
Om Kulkarni, Pierre-Emmanuel Sugier, Julie Guibon, Anne Boland-Augé, Christine Lonjou, Delphine Bacq-Daian, Robert Olaso, Carole Rubino, Vincent Souchard, Frédérique Rachedi, Juan Jesus Lence-Anta, Rosa Maria Ortiz, Constance Xhaard, Pierre Laurent-Puig,
Scientific Reports.2021;[Epub] CrossRef - Familial non medullary thyroid carcinoma: Beyond the syndromic forms
Aida Orois, Mireia Mora, Irene Halperin, Josep Oriola
Endocrinología, Diabetes y Nutrición (English ed.).2021; 68(4): 260. CrossRef - Discrimination between 34 of 36 Possible Combinations of Three C>T SNP Genotypes in the MGMT Promoter by High Resolution Melting Analysis Coupled with Pyrosequencing Using A Single Primer Set
Katja Zappe, Christine Pirker, Heidi Miedl, Martin Schreiber, Petra Heffeter, Georg Pfeiler, Stefan Hacker, Werner Haslik, Sabine Spiegl-Kreinecker, Margit Cichna-Markl
International Journal of Molecular Sciences.2021; 22(22): 12527. CrossRef - Emerging Biomarkers in Thyroid Practice and Research
Shipra Agarwal, Andrey Bychkov, Chan-Kwon Jung
Cancers.2021; 14(1): 204. CrossRef - The effects of the genetic polymorphisms of antioxidant enzymes on susceptibility to papillary thyroid carcinoma
Saeedeh Salimi, Mahdiyeh Harati‐Sadegh, Moein Eskandari, Zahra Heidari
IUBMB Life.2020; 72(5): 1045. CrossRef - Familial Aggregation and Heritability of Nonmedullary Thyroid Cancer in an Asian Population: A Nationwide Cohort Study
Huan-Tang Lin, Fu-Chao Liu, Shu-Fu Lin, Chang-Fu Kuo, Yu-Ying Chen, Huang-Ping Yu
The Journal of Clinical Endocrinology & Metabolism.2020; 105(7): e2521. CrossRef - Does the TT Variant of the rs966423 Polymorphism in DIRC3 Affect the Stage and Clinical Course of Papillary Thyroid Cancer?
Kinga Hińcza, Artur Kowalik, Iwona Pałyga, Agnieszka Walczyk, Danuta Gąsior-Perczak, Estera Mikina, Tomasz Trybek, Monika Szymonek, Klaudia Gadawska-Juszczyk, Klaudia Zajkowska, Agnieszka Suligowska, Artur Kuchareczko, Karol Krawczyk, Janusz Kopczyński, M
Cancers.2020; 12(2): 423. CrossRef - The effect of TP53 and P21 gene polymorphisms on papillary thyroid carcinoma susceptibility and clinical/pathological features
Zahra Heidari, Mahdiyeh Harati‐Sadegh, Abtin Arian, Rostam Maruei‐Milan, Saeedeh Salimi
IUBMB Life.2020; 72(5): 922. CrossRef - Feasibility Study Shows Multicenter, Observational Case-Control Study Is Practicable to Determine Risk of Secondary Breast Cancer in Females With Differentiated Thyroid Carcinoma Given Radioiodine Therapy in Their Childhood or Adolescence; Findings Also S
Valentina Drozd, Rita Schneider, Tamara Platonova, Galina Panasiuk, Tatjana Leonova, Nataliya Oculevich, Irina Shimanskaja, Irina Vershenya, Tatjana Dedovich, Tatjana Mitjukova, Inge Grelle, Johannes Biko, Christoph Reiners
Frontiers in Endocrinology.2020;[Epub] CrossRef - Genetic Mutations and Variants in the Susceptibility of Familial Non-Medullary Thyroid Cancer
Fabíola Yukiko Miasaki, Cesar Seigi Fuziwara, Gisah Amaral de Carvalho, Edna Teruko Kimura
Genes.2020; 11(11): 1364. CrossRef - Current Knowledge of Germline Genetic Risk Factors for the Development of Non-Medullary Thyroid Cancer
Kinga Hińcza, Artur Kowalik, Aldona Kowalska
Genes.2019; 10(7): 482. CrossRef - Thyroid Cancer: The Quest for Genetic Susceptibility Involving DNA Repair Genes
Santos, Gomes, Bastos, Gil, Azevedo, Ferreira, Limbert, Silva, Rueff
Genes.2019; 10(8): 586. CrossRef - Effect of the thymine‐DNA glycosylase rs4135050 variant on Saudi smoker population
Mikhlid Almutairi, Abdullah Mohammad Alhadeq, Rafa Almeer, Mohammed Almutairi, Mohammed Alzahrani, Abdelhabib Semlali
Molecular Genetics & Genomic Medicine.2019;[Epub] CrossRef - Association of Vitamin D Pathway Genetic Variation and Thyroid Cancer
Isabel S. Carvalho, Catarina I. Gonçalves, Joana T. Almeida, Teresa Azevedo, Teresa Martins, Fernando J. Rodrigues, Manuel C. Lemos
Genes.2019; 10(8): 572. CrossRef - Association of radiation risk in the second and third generations with polymorphisms in the genes CYP1A1, CYP2E1, GSTP1 and changes in the thyroid
Meruyert Massabayeva, Nailya Chaizhunusova, Nurlan Aukenov, Tolkyn Bulegenov, Bakytbek Apsalikov, Aigerim Shapihanova, Yersin Zhunussov
Molecular Medicine.2019;[Epub] CrossRef - BRAF‐positive multifocal and unifocal papillary thyroid cancer show different messenger RNA expressions
Kyoungjune Pak, Sunghwan Suh, Tae Sik Goh, Seong‐Jang Kim, Sae‐Ock Oh, Ju Won Seok, In Joo Kim, Yun Hak Kim
Clinical Endocrinology.2019; 90(4): 601. CrossRef

 KES
KES

 PubReader
PubReader ePub Link
ePub Link Cite
Cite